Classification de pixels dans des images multi-canaux#
APOC accepte également des listes d’images comme entrée pour l’entraînement et la prédiction. Cela peut être utilisé par exemple pour la segmentation sémantique dans des images multi-canaux montrant des noyaux, des membranes et du cytoplasme entre les deux.
from skimage.data import cells3d
from skimage.io import imsave, imread
import napari
import numpy as np
import matplotlib.pyplot as plt
from pyclesperanto_prototype import imshow
image = cells3d()
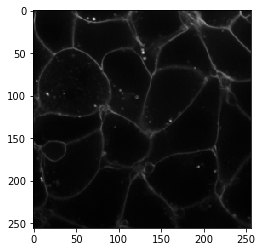
image_ch1 = image[30, 0]
imshow(image_ch1)
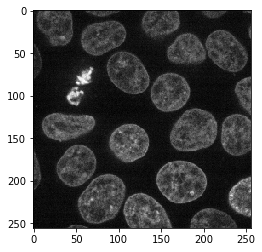
image_ch2 = image[30, 1]
imshow(image_ch2)
filename = '../../data/cells_annotation.tif'
annotation = imread(filename)
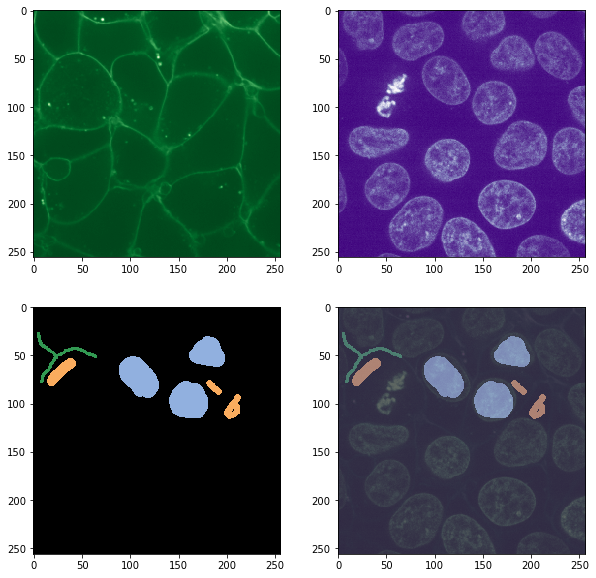
fix, axs = plt.subplots(2,2, figsize=(10,10))
imshow(image_ch1, plot=axs[0,0], colormap="Greens_r")
imshow(image_ch2, plot=axs[0,1], colormap="Purples_r")
imshow(annotation, labels=True, plot=axs[1,0])
imshow(image_ch1, continue_drawing=True, plot=axs[1,1], colormap="Greens_r", alpha=0.5)
imshow(image_ch2, continue_drawing=True, plot=axs[1,1], colormap="Purples_r", alpha=0.5)
imshow(annotation, labels=True, plot=axs[1,1], alpha=0.5)
Entraînement#
from apoc import PixelClassifier
# define features
features = "sobel_of_gaussian_blur=2 laplace_box_of_gaussian_blur=2 gaussian_blur=2 sobel_of_gaussian_blur=4 laplace_box_of_gaussian_blur=4 gaussian_blur=4"
# this is where the model will be saved
cl_filename = 'test.cl'
clf = PixelClassifier(opencl_filename=cl_filename)
clf.train(features=features, ground_truth=annotation, image=[image_ch1, image_ch2])
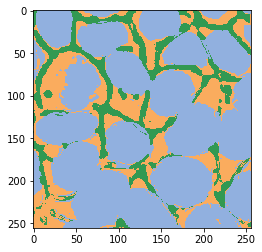
Prédiction#
result = clf.predict(image=[image_ch1, image_ch2])
imshow(result, labels=True)

